I study viruses: How our team isolated the new coronavirus to fight the global pandemic
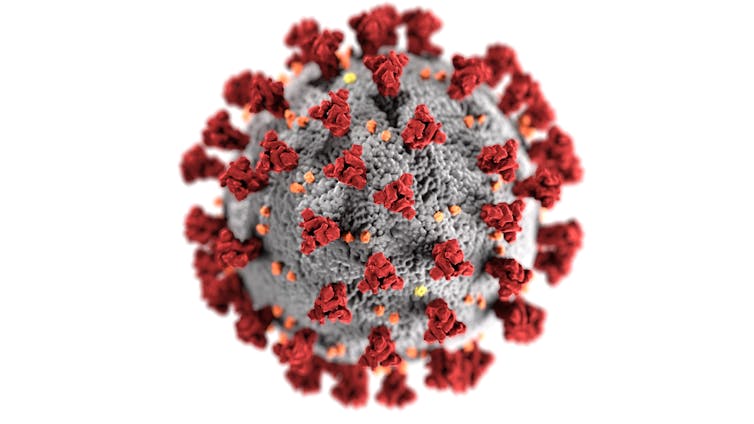
As most people rush to distance themselves from COVID-19, Canadian researchers have been waiting eagerly to get our (gloved) hands on the hated virus.
We want to learn everything we can about how it works, how it changes and how it interacts with the human immune system, so we can test drugs that may treat it, develop vaccines and diagnostics and prevent future pandemics.
This is what researchers live to do. Much of our everyday work is incremental. It’s important and it moves the field forward, but to have a chance to contribute to fighting a pandemic is especially inspiring and exciting.
The secret lives of viruses
Viruses are fascinating. They are inert microscopic entities that can either hide out, innocuous and undetected, or wreak pandemic havoc.
They are simultaneously complex and simplistic, which is what makes them so interesting — especially new, emerging viruses with unique characteristics. Researching viruses teaches us not only about the viruses we study, but also about our own immune systems.
Read more: Coronavirus weekly: expert analysis from The Conversation global network
The emergence of a new coronavirus in a market in Wuhan, China, in December 2019 set in motion the pandemic we are now witnessing in 160 countries around the world. In just three months, the virus has infected more than 360,000 people and killed more than 16,000.
Viral isolation
The outbreak sent researchers around the world racing to isolate laboratory specimens of the virus that causes COVID-19. The virus was later named severe acute respiratory syndrome coronavirus 2, or SARS-CoV-2.
In countries that experienced earlier outbreaks, including China, Australia, Germany and the United States, researchers were able to isolate the virus and develop their own inventories of SARS-CoV-2, but logistical and legal barriers prevented them from readily sharing their materials with researchers beyond their borders.
What Canadian researchers needed to join the fight in earnest was a domestic supply of clean copies of the virus — preferably from multiple Canadian COVID-19 cases. Even in a pandemic, developing such a supply is not as easy as it might sound, and multiple teams in Canada set out to isolate and develop pure cultures of the virus, not knowing which would be successful, or when.
Ultimately two teams in Canada would isolate the virus for study: one at the University of Saskatchewan and one that featured researchers from McMaster University, Sunnybrook Health Sciences Centre and the University of Toronto.
Arinjay Banerjee, a postdoctoral research fellow at McMaster who typically works in my virology lab, volunteered his special expertise. We were proud to have him share his talent with the team in Toronto, where he set to work with physicians and researchers Samira Mubareka, Lily Yip, Patryk Aftanas and Rob Kozak.
For Banerjee, it was like a batter being called to the plate with the score tied in the bottom of the ninth. He had come to work at McMaster because of its Institute for Infectious Disease Research and its Immunology Research Centre, and because the university maintains a research colony of bats.
Banerjee’s PhD work at the University of Saskatchewan, and now at McMaster, has focused on bats and how their viruses, including coronaviruses, interact with bat and human antiviral responses. Over the past few years, studies have shown that bat coronaviruses have the capacity to infect human cells. Multiple researchers had predicted a coronavirus that would evolve and jump into humans.
Read more: It's wrong to blame bats for the coronavirus epidemic
Ideal viral conditions
Isolating a virus requires collecting specimens from patients and culturing, or growing, any viruses that occur in the samples. These viruses are obligate intracellular parasites, which means that they can only replicate and multiply in cells. To isolate a particular virus, researchers need to provide it with an opportunity to infect live mammalian cells, in tiny flasks or on tissue culture plates.
Viruses adapt to their hosts and evolve to survive and replicate efficiently within their particular environment. When a new virus such as SARS-CoV-2 emerges, it isn’t obvious what particular environment that virus has adapted to, so it can be hard to grow it successfully in the lab.
We can use tricks to draw out a virus. Sometimes the tricks work and sometimes they don’t. In this case, the researchers tried a method Banerjee and the team had previously used while working on the coronavirus that causes Middle Eastern Respiratory Syndrome: culturing the virus on immunodeficient cells that would allow the virus to multiply unchecked. It worked.
Since specimens from patients are also likely to contain other viruses, it is critical to determine if a virus growing in the culture is really the target coronavirus. Researchers confirm the source of infection by extracting genetic material from the virus in culture and sequencing its genome.
They compare the sequence to known coronavirus sequences to identify it precisely. Once a culture is confirmed, researchers can make copies to share with colleagues.
All this work must be done in secure, high-containment laboratories that mitigate the risk of accidental virus release into the environment and also protect scientists from accidental exposure. The more versions of a virus that can be isolated, the better. Having multiple virus isolates allows us to monitor how the virus is evolving in humans as the pandemic progresses. It also allows researchers to test the efficacy of vaccines and drugs against multiple mutations of the virus.
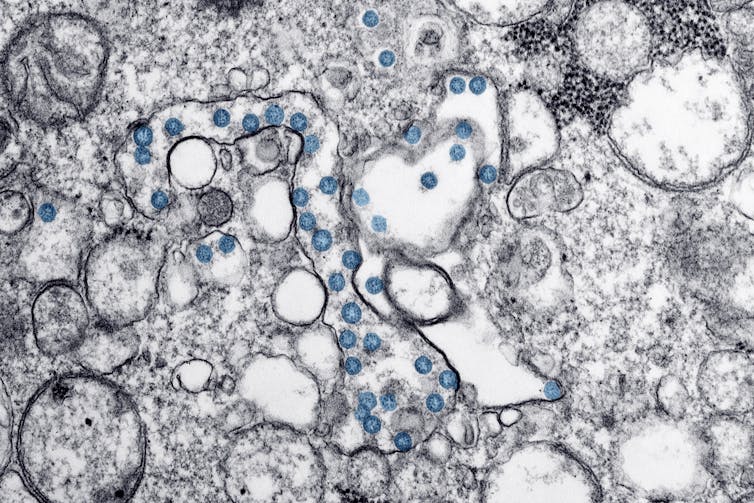
Canadian viral strains
Both the Saskatchewan and Ontario teams are now able to make and share research samples with other Canadian scientists, enabling important work to proceed, using a robust domestic supply that reflects the evolving virus in its most relevant mutations.
That in turn gives Canadian researchers a fighting chance to deliver a meaningful blow to COVID-19 while there is still time. I’m glad our colleagues at other Canadian institutions will also have versions of the virus to use in their research.
There is still so much work for all of us to do.
Karen Mossman, Professor of Pathology and Molecular Medicine and Acting Vice President, Research, McMaster University
This article is republished from The Conversation under a Creative Commons license. Read the original article.
.jpg.bee6da21545b36ee147b8b8fe68bc87e.jpg)
